Supercomputing, big data and artificial intelligence are crucial tools in the fight against the coronavirus pandemic. Around the world, researchers, corporations and governments are urgently devoting their computing resources to this global crisis. This column collects the biggest news about how advanced technologies are helping us fight back against COVID-19.
White House announces anti-COVID-19 supercomputing initiative
The White House announced a new public-private consortium to leverage U.S. supercomputing against the coronavirus. The COVID-19 High Performance Computing Consortium, led by the White House Office of Science and Technology Policy, “pools 16 systems that together offer over 330 petaflops of supercomputing capacity.” The release promises additional capacity in the future. “America is coming together to fight COVID-19, and that means unleashing the full capacity of our world-class supercomputers to rapidly advance scientific research for treatments and a vaccine. We thank the private sector and academic leaders who are joining the federal government as part of the Trump Administration’s whole-of-America response,” said Michael Kratsios, U.S. Chief Technology Officer. To read more, click here and here.
Frontera prepares for massive coronavirus simulations
The Frontera supercomputer at the Texas Advanced Computing Center (TACC) is working to take COVID-19 simulations to the next level. Once complete, the simulations will yield the first all-atom model of the coronavirus envelope. “If we have a good model for what the outside of the particle looks like and how it behaves, we’re going to get a good view of the different components that are involved in molecular recognition,” said Rommie Amaro, who is leading the efforts. So far, Frontera has helped optimize code and test the first parts of the model. To read more, click here.
Rosetta@home rallies volunteer computers against the coronavirus
Much like Folding@home, Rosetta@home is a network of crowdsourced computers that helps researchers examine protein-related computing problems. Measuring at around 1.26 petaflops of power, Rosetta@home (like Summit) crunched the crucial spike protein of the coronavirus, producing an atomic-scale structure of the protein weeks before its structure was determined in a lab setting. Now, Rosetta@home’s work is being used to develop vaccine trials and therapeutics for COVID-19. To read more about Rosetta@home’s work, visit the HPCwire article here.
Globus offers free access to COVID-19 researchers
Globus, a software-as-a-service company that assists with research data management for high-performance computing, has announced that, effectively immediately, they are “offering access to all Globus features at no cost to any institution engaged in COVID-19 research.” Globus stressed that this included features only available to subscribers and offered one-on-one consultations for COVID-19 researchers. To read more, click here.
 PRACE fast-tracks COVID-19 proposals
PRACE fast-tracks COVID-19 proposals
The Partnership for Advanced Computing in Europe (PRACE) has announced that it is welcoming and fast-tracking proposals that contribute to the “mitigation of the impact of the COVID-19 pandemic.” By way of example, they mention research to understand the mechanisms of infection, research to understand mutations and evolution, simulations to develop therapeutics or vaccines and analysis to understand the spread of the virus. To read more, click here.
GENCI offers its resources for coronavirus research
GENCI, the French national high-performance computing organization, has announced that it is providing its computing and storage resources to COVID-19 researchers. Like PRACE, GENCI highlights uses for vaccines and therapies, epidemiological research on the spread of the virus and related applications. To read more, click here.
BioTeam, an interdisciplinary group of experts that offers computing services for life science applications, is offering up to 40 hours of services free of charge to COVID-19 researchers. The allocation (available for “scientific computing, cloud, IT and strategic consulting services”) is accessible for laboratories and data science groups directly developing therapies or vaccines for COVID-19. Furthermore, BioTeam announced that it was offering its own computational power to the Folding@home project, which is working on coronavirus problems (to read more on that, visit the HPCwire article). To read more about BioTeam, click here.
Rensselaer lends its supercomputing to the COVID-19 fight
Rensselaer Polytechnic Institute has announced that AiMOS, currently the 24th most powerful publicly ranked supercomputer, will be available for use by researchers working against COVID-19. The institute is reaching out to the research community to offer access. “This effort requires expertise, collaboration, and the ability to process incredible amounts of data, and Rensselaer is offering all three at this critical time,” said Shirley Ann Jackson, president of Rensselaer. “In particular, the ability to model at very large scales requires the unique capabilities of AiMOS.” To read more, click here.
Singapore’s National Supercomputing Centre turns its eye to the coronavirus
The National Supercomputing Centre (NSCC) in Singapore has offered its resources to Singapore scientists who want to study COVID-19, issuing a special call for projects that extends until September 2020. Proposals, they say, will be fast-tracked and, if approved, offered priority access to resources, which include the ASPIRE 1 petascale supercomputer. The applications will be reviewed by a special COVID-19 selection committee composed of bioinformatics and molecular biology experts. To read more, click here.
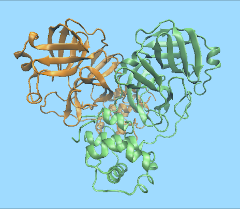
RIKEN explores COVID-19’s molecular dynamics
Researchers at the RIKEN Center for Biosystems Dynamics in Japan have used a drug research supercomputer to analyze the structure of the coronavirus. Specifically, they examined COVID-19’s main protease, which is one of the mechanisms the virus uses for self-replication. The 10-microsecond-long simulation took 10 days on the MDGRAPE-4A supercomputer, and the researchers have made the results available free of charge. To read more, click here.
Do you know about COVID-19 research that should be featured on this list? If so, send us an email at [email protected]. We look forward to hearing from you.





























































