Supercomputing, big data and artificial intelligence are crucial tools in the fight against the coronavirus pandemic. Around the world, researchers, corporations and governments are urgently devoting their computing resources to this global crisis. This column collects the biggest news about how advanced technologies are helping us fight back against COVID-19.
AWS highlights its many COVID-19 collaborations
Amazon Web Services (AWS) highlighted a slew of AWS research collaborations targeting COVID-19. These projects range from helping PostEra find an antiviral cure for COVID-19 within four months to helping Emory University use text mining on the scientific literature surrounding COVID-19. To read more about AWS’ collaborations, click here.
 Lawrence Livermore introduces new COVID-19 research portal
Lawrence Livermore introduces new COVID-19 research portal
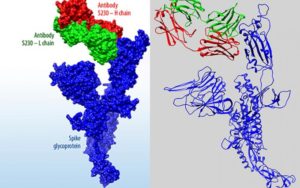
Lawrence Livermore National Laboratory (LLNL) has introduced a new, searchable data portal through which it will share its COVID-19 research with scientists and the general public. Of particular interest here are the results of LLNL’s virtual screenings of small molecules and designed antibodies, which were tested on their ability to bind to COVID-19. To read more about the research portal, click here.
The Watson AI Lab funds ten projects addressing COVID-19
The MIT-IBM Watson AI Lab has funded ten MIT-based projects aiming to use AI to confront the pandemic. These projects include early detection of sepsis in COVID-19 patients; protein design for hindering COVID-19; optimization techniques for designing lockdown strategies; face mask material selection; and more. To read more about these projects, click here.
 TACC supercomputers delve into COVID-19’s spike protein
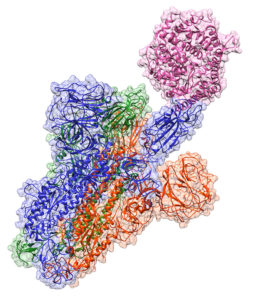
TACC supercomputers delve into COVID-19’s spike protein
Researchers from the University of Texas at El Paso (UTEP) are using supercomputers at the Texas Advanced Computing Center (TACC) to build complex models showing how COVID-19’s spike protein interacts with the human ACE2 protein – the process through which COVID-19 infects the human body. The researchers used the Stampede2 and Lonestar 5 supercomputers. To read more, click here.
MolSSI launches an open source repository for COVID-19 drug simulation data
The Molecular Sciences Software Institute (MolSSI) has launched an open-source website allowing biomolecular scientists to share their drug testing simulations targeting COVID-19’s viral proteins. “This repository, and the data within, are designed to get the information out quickly. Researchers really wanted to get their data out in front of other scientists even though it had not necessarily been through the peer review process yet,” said T. Daniel Crawford, lead director of the institute. To read more, click here.
Pawsey accelerates genome assembly for early COVID-19 detection
A team of researchers used resources at the Australian Pawsey Supercomputing Centre to develop a “fast, sensitive diagnostic test” for COVID-19. According to Pawsey, the new test can reliably detect lower concentrations of the virus and provide more genetic information about the virus. The test, called the “Pathogen-Oriented Low-cost Assembly and Resequencing” (or “POLAR”) test, is seeking FDA approval. To read more, click here.
Lab behind the ‘cloud burst’ joins Folding@home’s COVID-19 research
The Wisconsin IceCube Particle Astrophysics Center (WIPAC), which helped to orchestrate the record-breaking GPU “cloud burst” last year, is now dedicating a large portion of its on-premises computing resources to the crowdsourced Folding@home initiative. “It just feels right to make the effort to share computing resources from fields as far removed from virology as neutrino astrophysics,” says Kael Hanson, WIPAC’s director. “We’re pleased to aid in research that could ultimately lead everyone impacted by the current COVID-19 situation out of the crisis.” To read more, click here.
Do you know about COVID-19 research that should be featured on this list? If so, send us an email at [email protected]. We look forward to hearing from you.





























































