A research effort built around an automation platform that connects data and analytics applications will focus on advancing the use of AI and machine learning in bioscience and medical applications.
The centerpiece of the R&D agreement between ICM University of Warsaw and Tokyo-based Systems Biology Institute (SBI) is the Garuda connectivity and automation platform. Garuda is used to link devices and tools ranging from open-source Jupyter notebooks that share code and visualizations to bioscience apps.
The partners said Monday (Dec. 14) they would initially focus on AI-driven bioinformatics and genomics applications as well as computational systems biology.
Longer term, SBI is seeking the push the frontiers of AI research using tools like Garuda as platforms to develop a computational engine capable of autonomously making scientific discoveries worthy of the Nobel Prize.
SBI’s current research focuses on using AI technologies that advance bioscience research and applications for healthcare, medicine and climate science. Along with connecting researchers to HPC resources via Garuda, the collaboration with Polish researchers will use a text analysis platform called Taxila and a machine learning tool called Gandhara.
The joint research effort also will use Garuda for “-omics” data science applications, a reference to genomics and other bioscience disciplines that increasingly rely on HPC platforms, data analytics and AI tools. The products of the technology framework include new data pipelines and other tools promoted as platform agnostic.
The collaboration promises to jumpstart bioscience research in central Europe in disciples such as bioinformatics and biomedical data science, said Marek Michalewicz, director of the Interdisciplinary Center for Mathematical and Computational Modelling at the University of Warsaw. Garuda represents “the gold standard [for] how certain solutions should be developed and presented to users,” Michalewicz said in announcing the collaboration with SBI.
“We also plan to incorporate Garuda as one of the best multi-method, multi-tool computational solutions in our new multi-cloud infrastructure,” he added.
The partners added they would collaborate on AI and other software applications tailored to biomedical research. Those efforts would use Garuda as “connectivity layer for HPC,” said Hiroaki Kitano, president of the Systems Biology Institute.
Collaboration with the University of Warsaw builds on the Institute’s previous technology partnerships, including with Singapore’s Agency for Science, Technology and Research.
Katano, who also serves as president and CEO of Sony Computer Science Laboratory, has advocated development of AI systems that eventually contribute Nobel Prize-worthy scientific discoveries in physiology and medicine. The initiative is part of a Nobel Turing Challenge, which seeks to push the frontiers of AI research in areas like unsupervised learning and what Katano calls “machine creativity.”
Kitano framed the challenge this way: “Can we build machines that can win the Nobel Prize?”
That would entail convincing the “Nobel Committee to give [an] AI system the Nobel Prize without noticing it is an AI system, not a human scientist,” Kitano added.
The first step involves developing AI platforms “that can perform biomedical and biotechnology research fully autonomously that leads to major discoveries.” Then, the machine would have to be able to explain its reasoning behind a discovery, along with potential applications and social implications—otherwise known as explainable AI.
“What we are trying to do is use the computing power of AI to overcome and complement limitations on human cognition,” creating what Kitano promotes as an “engine of scientific discovery.” Such autonomous systems would for example be able to design and perform science experiments, “redefining scientific discovery,” Kitano asserted.
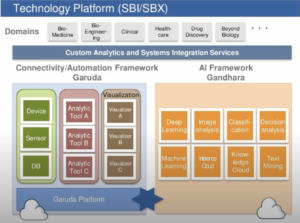
The technology platform needed to achieve that vision is Garuda, which is used to control computing assets and connect data analysis and simulation tools and components, including sensors and other monitoring devices. The AI stack includes the Gandhara framework that enables the platform to use AI modules.
In one example, Kitano said Japanese researchers are using Garuda for physiological, cell and ion-channel modeling using data assets that include a machine learning model database, medical imagery and genomic data. The modeling of entire organs can then be used in diagnostic applications.
Garuda is also being deployed in drug discovery efforts that combine machine learning models with molecular integration maps used to graph biological networks. Those efforts would be welcome since early efforts using AI tools in drug discovery have so far fallen far short of expectations.