Life sciences computational research has long been dominated by statistics and parallel processing more than traditional HPC. Think gene sequencing and variant calling. Mechanistic simulation and modeling has played a much smaller role though that’s changing. An exception is the Anton line of supercomputers, designed and built by D.E. Shaw Research (DESRES); these are purpose-built systems specifically for molecular dynamics modeling. At Hot Chips last week DESRES pulled back the covers on Anton 3 – which the design team dubbed a “Fire-Breathing Monster for Molecular Dynamics Simulations.”
 Not only is Anton 3 a major advance over Anton 2 but it’s an interesting departure from the current trend of packing supercomputers with blended AI and tightly-coupled HPC compute capability. Anton is aimed squarely at the latter, digging out the mechanistic details of molecular systems to unravel complicated biology and develop better drugs and therapy. Latency is the big challenge. A powerful ASIC with specialized cores to calculate electrostatic forces and bond energies is DESRES’s solution, along with streamlined communications.
Not only is Anton 3 a major advance over Anton 2 but it’s an interesting departure from the current trend of packing supercomputers with blended AI and tightly-coupled HPC compute capability. Anton is aimed squarely at the latter, digging out the mechanistic details of molecular systems to unravel complicated biology and develop better drugs and therapy. Latency is the big challenge. A powerful ASIC with specialized cores to calculate electrostatic forces and bond energies is DESRES’s solution, along with streamlined communications.
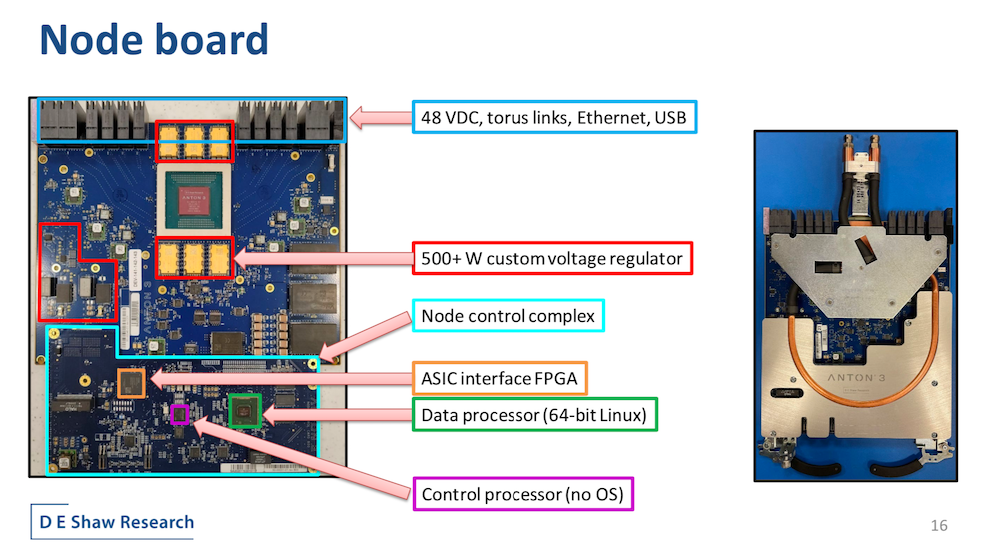
The approach, though not inexpensive, has worked well. Earlier this summer, DESRES completed the first “full-size 512-node” Anton 3 machine. It spans four racks, is water-cooled – no surprise – and the basic topology is a 3D torus. For the task at hand, it’s tough to beat.
“Anton 3’s highest performing competitor is an Nvidia A100, a relatively late model GPU,” said Adam Butts of DESRES in his talk. “Running Desmond – our in-house optimized MD code – on a chip for chip basis, Anton 3 is about 20 times faster than A100. This is not to pick on Nvidia. The A100 is a tremendously capable compute engine. Rather this illustrates the tremendous benefit we achieve from specialization.”
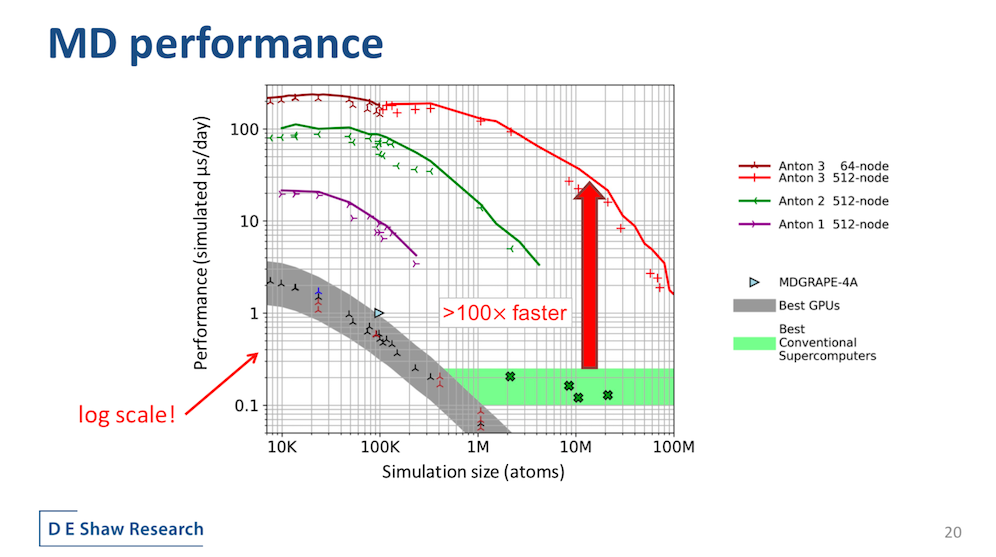
The metrics of merit for molecular simulation, generally, are how much real-time interaction of a molecular system can you simulate and how big a system of molecules can you simulate.
“Anton 3 represents the first Anton machine for which a single node is likely to be a useful size for scientific work. For example, at 113,000 atoms, the simulation of ACE2 (protein and target of CoV2 spike) that I showed early in the presentation fits quite comfortably on a single node Anton 3, turning in a performance more than an order of magnitude better than possible on a GPU,” said Butts.
“A distinguishing feature of Anton is that [performance] does scale with the higher node counts. Anton 3’s peak performance, below about 100,000 atoms, exceeds 200 microseconds per day using just 64 nodes. Thus, millisecond scale simulations are possible inside a workweek. The full-size 512-node machine maintains over 100 microseconds a day out to simulation sizes larger than 1 million atoms and supports simulations beyond 50 million atoms,” said Butts.
He added that, at least for now, using multiple GPUs to scale doesn’t help. “Until systems become very large, splitting a simulation across multiple GPUs does not currently yield enough benefit to repay the cost of internode communication.”
For many in HPC, the DESRES origin story is familiar. David E Shaw, a Columbia University professor turned hedge fund manager, was an early ‘quant’ and his spectacular success on Wall Street allowed him to turn his talents to biomedical research. He founded D.E. Shaw Research in 2001 with himself as chief scientist. This eventually led to Anton 1 (~2008) – the first purpose built supercomputer for molecular modeling and the Desmond software package for high-speed MD simulation. This was followed by Anton 2 (~2014), “the first platform to achieve simulation rates of multiple microseconds of physical time per day for systems with millions of atoms (IEEE abstract).”
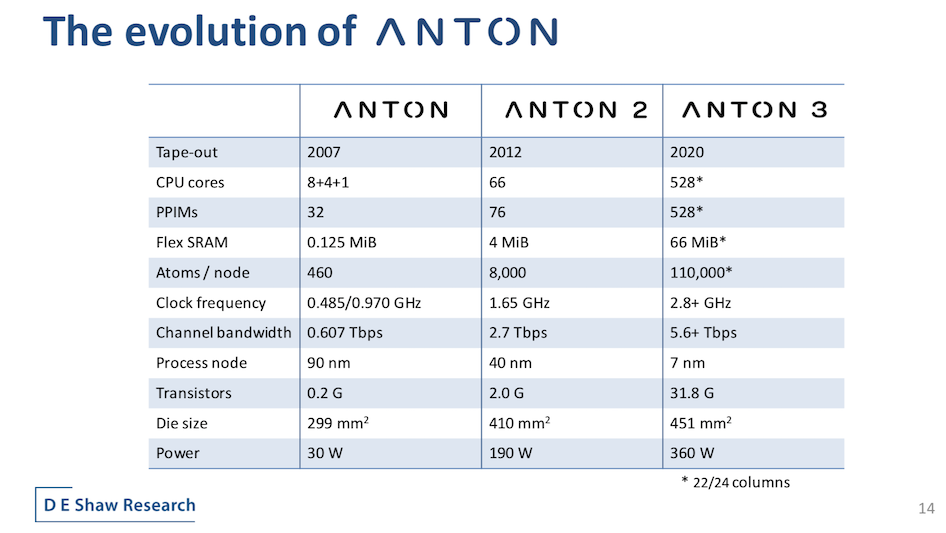
At the heart of the Anton line is its ASIC engine, which has undergone both incremental advance and dramatic innovation with each generation. As shown below (slide below) Anton’s capacity has steadily grown. Butt’s talk examined the Anton 2 ASIC’s architecture and then presented changes made to improve the Anton 3 chip. DESRES also has a paper[i] (Anton 3: Twenty Microseconds of Molecular Dynamics Simulation Before Lunch) scheduled for SC21. Presumably there will also be an SC21 talk.
Simulating large molecular systems, of course, is a difficult computational challenge. That’s why approximation techniques are so often used. Not with Anton, explained Butts, “Forces among 1000s to millions of atoms are computed according to physics-based models that describe the inter-atomic energy potentials. These forces are integrated over discrete time-steps of just a few femtoseconds to determine new positions and velocities for the atoms – a process repeated billions of times to generate trajectories at timescales of interest.”
It’s interesting to hear Butts describe the DESRES/Anton approach.
“We’ll start with a familiar electrostatic force between charged particles. Atoms within the simulation are assigned charges, which are parameters of the models, then the force between any pair of particles is proportional to the product of their charges divided by the square of their separation. Computing the interactions among all pairs of particles scales poorly, so the total force is rewritten as a sum of explicit pairwise interactions out to a cut-off radius, plus a distant contribution from charges beyond. That latter contribution can be expressed as a convolution which may be computed efficiently on a grid by multiplication in Fourier space,” he said.
“Of course, there’s more to the force field than the electrostatic interaction. Quantum mechanical effects are approximated through forces representing the topological connections implied by chemical bonds, and the Vander Waals force which acts between pairs of atoms but falls off quickly enough with distance that a range-limited computation is sufficiently accurate versus solving the underlying quantum mechanical equations. Such a model reduces the problem of long timescales simulation from intractable to merely ridiculous,” said Butts.
How Anton Breathes Fire
The Anton systems rely on specialized ASICs for their power. The Anton 2 ASIC was largely made of up of two computation tiles explained Butts (see slide below). “One – the flex tile – that looks like a typical multicore processor consisting of four cores with private caches connected to a shared memory. We call these geometry cores (GCs) due to their special facility for the vector geometric computations occurring frequently in MD code,” he said. A dispatch unit handles fine grained synchronization and the network interface connects the flex tile to the rest of the chip and machine.
The other called high throughput interaction subsystem or HTIS tile “is dominated by an array of pairwise point interaction modules or PPIMs. Each PPIM contains a pair of unrolled arithmetic pipelines, the PPIPs, which compute forces between pairs of interacting atoms. The PPIPs also participate in the distant force computation by spreading charges on the grid points and interpolating grid forces back onto atoms.”
“The interaction control block or ICB organizes the streaming of positions into the PPIM array and the unloading of the accumulated forces back out. Finally, a miniature version of a Flex tile performs command and control functions. The periphery of the chip contains the SerDes channels that interconnect the Anton 2 ASICs, the IO interfaces for connections to a host machine, and an on-chip logic analyzer,” said Butts.
Butts ticked through the major Anton 3 objectives: “We also have to address the performance bottlenecks exposed by accelerating the force computations. Computing bonds and GC code is one such bottleneck and the limited scaling of off-chip communication bandwidth is another. Besides making the machine faster, we also wanted to increase its capability, supporting larger simulations, making it easier to program and support new force field features within arithmetic pipelines. Finally, given that our design team (40ish people) is definitely not scaling according to Moore’s law, we need to control the complexity of the design and the implementation.”
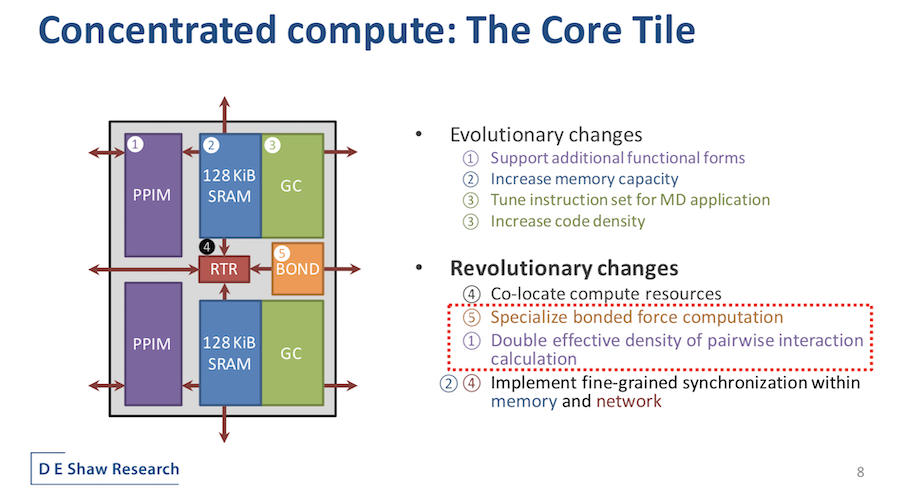
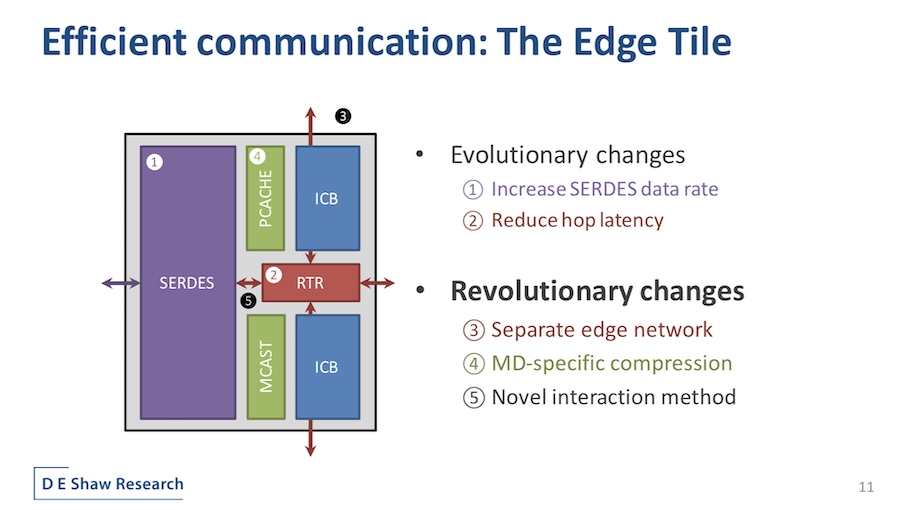
The new Anton 3 core tile, said Butts, distills all of the main components required for the MD computation into a handful of unique blocks. A central router provides on-chip network connections among a large array of such tiles. The GC and the PPIMs are more familiar from Anton 2 but underwent important evolution. The PPIMS, for example, implements new functional forms to support a broader range of force fields. Memory capacity per GC was doubled, “enabling larger simulations and more flexible software.” The GCs’ instruction set was also optimized with new instructions and denser encoding to increase the effective capacity of the GCs’ instruction cache.
“Besides these evolutionary changes, more significant changes have also occurred here, not least of which is the colocation of flexible and specialized compute resources into the same tile. Anton 3 supports bi-directional communication between the GCs and PPIMs, allowing for new use models involving fine-grained cooperation of the PPIMs’ high-throughput pair selection and the GCs’ programmability,” explained Butts. “The bond computation bottleneck has been relieved by introducing a new specialized pipeline for bonded forced computation. Pairwise interaction throughput per unit area is doubled thanks to a novel decomposition of the range-limited interactions. Finally, synchronization functionality is now distributed between the memory and network, eliminating the need for a separate dispatch unit.”
The Anton 3 ASIC’s clock is 2.8 gigahertz which helps both throughput and latency while raw channel bandwidth is more than doubled (Anton 2 1as 1.5 gigahertz). Both of these important parameters, said Butts, leave room for further improvement. Transistor count jumped 16x supported by the new 7nm FinFET process. Die size increase relative to Anton 2 was modest. Lastly, “ASIC power dissipation is just under twice that of Anton 2 to at 360 watts, although it is almost identical and normalized for die area and frequency,” said Butts.
Given this was a Hot Chips presentation, it was natural to focus on the chip’s details. Butts walked through innovations in power distribution and dug more deeply into a few areas such bond calculation improvements. He said little about Anton 3’s early research agenda but did note the early silicon went live without a hitch and was scaled up to 512 nodes relatively easily.
The big result of the upgrade is being able to simulate larger systems for longer times, which is key to making such granular MD simulation useful. Anton 3 should make doing that practical. Thus far there seems to be limited use of ML/DL in Anton though that could change.
Asked about data re-use and incorporation of more speculative AI into Anton, Brannon Batson a DESRES engineer who handled the post-talk Q&A said, “It’s very difficult in molecular dynamics to use stale data. Because of an important property of time-integrable systems, we need our simulations to only consume data that is current, it can’t be, you know, from previous time steps. You can maybe use that speculatively, if you have the ability to rollback, but I think the answer for us, at least thus far, is no, or if so we haven’t figured out how to do it yet.”
“Our computation is broadly divided into two classes, there’s kind of a more classical general purpose flexible subsystem. There we rely heavily on 32-bit fixed point operations in vector forms of those. But in our specialized arithmetic pipelines, we’re all over the place. There are places where we have 14-bit mantises with five-bit exponents, [and] there are places where we’re in log domain. Since we’re doing unrolled pipelines, we trim the precisions very carefully stage by stage basis, based on a very careful numeric analysis, and that’s where a lot of our value proposition comes from, for computational density,” he said.
Two interesting questions came from Jeffrey Vetter at Oak Ridge National Labs, who asked if the system could scale beyond 512 nodes for larger MD simulations, and whether other types of molecular dynamics applications were being considered, such as those focused on material science.
Batson said, “I can tell you that the hardware is physically capable of scaling larger than 512 nodes in terms of the network in the link layer. But the machine is definitely designed to operate normally for where a single molecular dynamic simulation runs on at most 512 nodes. The larger configurations would be if you want to run multiple simulations and sort of exchange data between them. There’s some facility for doing that.
“For the other applications, in particular material science, it’s possible there are applications that would benefit from Anton. We’ve looked at this somewhat within DESRES, but not a whole lot. Like I said before, you know, we’re not a computer company, our focus is on curing diseases and easing human misery and we just don’t spend a lot of time working on things outside of that scope.”
There’s always been a mixture of admiration and a little envy within the life sciences research community regarding DESRES’s Aton systems. Such systems are powerful but rarer.

Ari Berman, CEO of BioTeam, a life sciences computational research consultancy, noted, “The Anton systems have always tipped the balance of competitive edge in the field and they are always the holy grail of researchers solving complex biomolecular systems. Anton 3 looks to take that edge to an entirely new level, putting access to the system at a 100X advantage compared to even the best general use supercomputers available to researchers. These types of simulations are of critical importance since they cut years off of pharmaceutical development, help narrow the set of variables in understanding biological systems (such as disease-causing mutations) and a host of other applications.
“The most recent and powerful example is the effect structural biology and MD simulations had on our ability to create vaccines for COVID-19 at an unheard-of speed by quickly solving and resolving the mechanisms of the spike protein on the SARS-CoV-2 viral surface. However, there is only one Anton 3, and if you’d like to use it you’ll need to appeal to D.E. Shaw until and unless they make the system more widely available (like they did with Anton 2 at PSC in 2017). Perhaps this time around, the public clouds will take an interest in the technology and will work with D.E. Shaw to make the technology at least moderately more available to the community.”
Perhaps the SC21 paper (and presentation if there is one) will talk more about use cases for the latest Anton system.
[i] D. E. Shaw et al., “Anton 3: Twenty Microseconds of Molecular Dynamics Simulation Before Lunch,” to appear in SC’21: Proceedings of the International Conference for High Performance Computing, Networking, Storage, and Analysis, 2021.