In this regular feature, HPCwire highlights newly published research in the high-performance computing community and related domains. From parallel programming to exascale to quantum computing, the details are here.
 ScriptManager: an interactive platform for reducing barriers to genomics analysis
ScriptManager: an interactive platform for reducing barriers to genomics analysis
Researchers from Cornell University introduce ScriptManager, built to “take optimal advantage of the available compute resources, enabling a life sciences researcher to conduct scalable genomic analysis without requiring them to know the underlying computing systems.” According to the research team, the software includes a graphical interface that enables a user-friendly navigation “of inputs and options while also supporting a command line interface for automation and integration with workflow managers like Galaxy.” In their paper, the researchers describe how users with little experience using the command line, “can leverage national supercomputing resources using a graphical desktop interface like Open OnDemand.” The goal is to provide a tool in the genomics community that streamlines access and avoids technical difficulties facilitating the scientists’ workflows to be integrated into large-scale production pipelines. The source code is publically available at https://github.com/CEGRcode/scriptmanager.
Authors: Olivia W. Lang, Franklin Pugh, and William Km Lai
The OpenMP cluster programming model
In this paper, Brazilian researchers present OpenMP Cluster (OMPC), which is a “task-parallel model that extends OpenMP for cluster programming.” According to researchers from the University of Campinas, the Federal University of ABC, and Petrobras, a state-owned Brazilian company in the petroleum industry, “OMPC allows for the offloading of complex scientific tasks across HPC cluster nodes in a transparent and balanced way.” Currently, “experimental results show that OMPC can deliver up to 1.53x and 2.43x better performance than Charm++ on CCR and scalability experiments, respectively.” In addition, researchers demonstrated through experiments that “OMPC performance weakly scales for both Task Bench and a real-world seismic imaging application.”
Authors: Hervé Yviquel, Marcio Pereira, Emílio Francesquini, Guilherme Valarini, Gustavo Leite, Pedro Rosso, Rodrigo Ceccato, Carla Cusihualpa, Vitoria Dias, Sandro Rigo, Alan Sousa, and Guido Araujo
 Coupling streaming AI and HPC ensembles to achieve 100-1000× faster biomolecular simulations
Coupling streaming AI and HPC ensembles to achieve 100-1000× faster biomolecular simulations
A multidisciplinary team of researchers from Argonne National Laboratory, University of Illinois Urbana-Champaign, Rutgers University, and Brookhaven National Laboratory, discuss DeepDriveMD in this paper. DeepDriveMD is “a framework for ML-driven steering of scientific simulations that we have used to achieve orders-of-magnitude improvements in molecular dynamics (MD) performance via effective coupling of ML and HPC on large parallel computers.” Researchers used “three biophysical MD modeling applications to evaluate its design, implementation, and performance, and demonstrate that by driving ensembles of MD simulations with ML approaches.” The results “demonstrate that DeepDriveMD can achieve between 100–1000× acceleration for protein folding simulations relative to other methods, as measured by the amount of simulated time performed, while covering the same conformational landscape as quantified by the states sampled during a simulation.”
Authors: Alexander Brace, Igor Yakushin, Heng Ma, Anda Trifan, Todd Munson, Ian Foster, Arvind Ramanathan, Hyungro Lee, Matteo Turilli, and Shantenu Jha
Quantum computational advantage with a programmable photonic processor
Researchers from the National Institute of Standards and Technology detail their experiences achieving “quantum computational advantage using Borealis, a photonic processor offering dynamic programmability on all gates implemented.” Researchers “carry out Gaussian boson sampling (GBS) on 216 squeezed modes entangled with three-dimensional connectivity, using a time-multiplexed and photon-number-resolving architecture.” Using Borealis, the authors claim “it would take more than 9,000 years for the best available algorithms and supercomputers to produce, using exact methods, a single sample from the programmed distribution, whereas Borealis requires only 36 μs.”
Authors: Lars S. Madsen, Fabian Laudenbach, Mohsen Falamarzi. Askarani, Fabien Rortais, Trevor Vincent, Jacob F. F. Bulmer, Filippo M. Miatto, Leonhard Neuhaus, Lukas G. Helt, Matthew J. Collins, Adriana E. Lita, Thomas Gerrits, Sae Woo Nam, Varun D. Vaidya, Matteo Menotti, Ish Dhand, Zachary Vernon, Nicolás Quesada, and Jonathan Lavoie
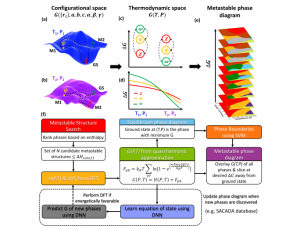
Machine learning the metastable phase diagram of covalently bonded carbon
A multidisciplinary team of researchers from Argonne National Laboratory, University of Illinois, and Northwestern Argonne Institute of Science and Engineering in Illinois, and the Center for High Pressure Science and Technology Advanced Research in China “introduce an automated workflow that integrates first-principles physics and atomistic simulations with machine learning and high-performance computing.” This workflow will “allow rapid exploration of the metastable phases to construct ‘metastable’ phase diagrams for materials far-from-equilibrium.” The researchers used carbon as a prototypical system and demonstrated “automated metastable phase diagram construction to map hundreds of metastable states ranging from near equilibrium to farfrom-equilibrium (400 meV/atom).”
Authors: Srilok Srinivasan, Rohit Batra, Duan Luo, Troy Loeffler, Sukriti Manna, Henry Chan, Liuxiang Yang, Wenge Yang, Jianguo Wen, Pierre Darancet, and Subramanian K.R.S. Sankaranarayanan
Distributed-memory parallel contig generation for de novo long-read genome assembly
A research team with affiliations at University of California, Berkeley, and Lawrence Berkeley National Laboratory tackle the computing challenges associated with de novo genome assembly, which is a strategy used to rebuild a “sequence of an unknown genome from redundant and erroneous short sequences.” With their paper, the researchers sought to “build on previous work in the literature, diBELLA 2D, and present a novel distributed-memory algorithm that generates the contig set starting from a string graph representation of the genome and using a sparse matrix abstraction.” Using the Haswell partition of the Cray XC40 supercomputer Cori at NERSC and the IBM supercomputer Summit at Oak Ridge National Laboratory, the researchers evaluated their contig generation algorithms and the ELBA assembly pipeline. The algorithm created by the researchers demonstrated a “good scaling with parallel efficiency up to 80% on 128 nodes, resulting in uniform genome coverage and showing promising results in terms of assembly quality.” The “algorithm localizes the assembly process to significantly reduce the amount of computation spent on this step.”
Authors: Giulia Guidi, Gabriel Raulet, Daniel Rokhsar, Leonid Oliker, Katherine Yelick, and Aydin Buluc
Asynchronous distributed Bayesian optimization at HPC scale
A team of researchers from the Argonne National Laboratory (U.S.), INP-ENSEEIHT (France) and Universite Paris-Saclay (France), tackle “the computational overhead in multipoint generation schemes, [considered] a major bottleneck in designing BO methods that can scale to thousands of workers.” In their paper, the researchers develop an “asynchronous-distributed BO method wherein each worker runs a search and asynchronously communicates the input-output values of black-box evaluations from all other workers without the manager.” The researchers scale their “method up to 4,096 workers and demonstrate improvement in the quality of the solution and faster convergence.” In addition, they were able to show the effectiveness of their “approach for tuning the hyperparameters of neural networks from the Exascale Computing Project CANDLE benchmarks.”
Authors: Romain Egele, Joceran Gouneau, Venkatram Vishwanath, Isabelle Guyon, and Prasanna Balaprakash
Do you know about research that should be included in next month’s list? If so, send us an email at [email protected]. We look forward to hearing from you.



























































