Artificial-intelligence algorithm developed on XSEDE-allocated systems promises better MRI agents with unprecedented speed of discovery
Jan. 6, 2022 — Using XSEDE-allocated supercomputers, scientists at Carnegie Mellon University (CMU) have created an “artificial chemist,” a computer program that mimics the expertise of human chemists. The artificial intelligence (AI) system is capable of directing an automated laboratory to synthesize new contrast agents for medical MRI imaging. The new contrast agents, thanks to the AI, have a ratio of signal-to-noise as much as 50% higher than previous state-of-the-art, human-designed materials. This performance boost offers the possibility of more detailed medical scans of the human body, improving diagnosis.
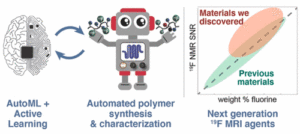
The CMU team, led by prof. Olexandr Isayev, built their AI using advanced research computers at XSEDE resources the Pittsburgh Supercomputing Center (PSC) and the Texas Advanced Computing Center (TACC). The robotic lab instrument is located at the University of North Carolina at Chapel Hill (UNC). The collaborators plan to develop the software so that it’s capable of more general chemical design for other applications in medicine, chemistry and materials science.
Why It’s Important
New chemical compounds – particularly a type of chemical called a polymer, which is made up of smaller building blocks called monomers – are a mainstay of advancement in chemistry, medicine, computing and other fields. A major limitation in this field, though, is that humans, who learn from each other, may be in “ruts,” working the same way they always have and not seeing promising alternatives. The CMU-led team wanted to create a general-purpose AI chemist that could teach itself how to select combinations of monomers, avoiding human bias.
“Previous efforts in materials discovery have relied on either luck or human intuition, which both suffer from inherent biases and limitations in knowledge,” said Olexandr Isayev at CMU.
To create an AI chemist that could forge new paths in synthetic chemistry, the scientists used an approach called automated machine learning. The first step in the process is the usual one of selecting the best performer from a large group of possible machine learning models. The scientists then refine the winning model by real-world testing of the resulting contrast-agent candidates in the UNC lab, putting the results of that testing back into the AI. By going back and forth between the computer and the lab, the AI could correct its mistakes and biases.
The overall project posed serious challenges. The first was a truly vast number of possible polymers that could be produced by the automated lab’s reagent set. Unlike working with simulated or historical data, the development of an AI algorithm plus acquiring new data on the fly by real experiments required taking into account the cost and number of experiments. To some extent, the team could control this by giving the AI a limited set of reagents – but even for a small set of six organic monomers, the space of possible experiments was over 50,000. The CMU team would need powerful computing resources. They would also have to refine the AI model in repeated training steps to cover the huge multidimensionality of the problem in search of the best-performing polymers while conducting only a small fraction of possible experiments. To overcome this challenge, they turned to XSEDE.
How XSEDE Helped
XSEDE supplied the group with access to powerful supercomputers containing graphics processing units, or GPUs. These processors were originally designed to create realistic images for computer gaming. But their unique capabilities for “parallel computing” proved to be ideal for AI research. Starting in 2012, a GPU revolution swept the AI field, powering many of the groundbreaking AI tools we now take for granted.
PSC’s XSEDE-allocated Bridges-2 system, as well as the NSF Petascale Computing Resource Frontera at TACC, offered the team powerful new GPUs to “train” the AI, as well as the massive memory (RAM) needed to keep track of the problem’s multidimensionality.
Filipp Gusev, a graduate student in Isayev’s group in the joint CMU-University of Pittsburgh PhD Program in Computational Biology, designed the AI software in a way that didn’t rely on historically biased knowledge as experienced by humans. Instead, in a process called machine learning, or ML, it started with a small “training set” of successful MRI contrast agents to use as a starting point for the model. By acquiring new information in a process called active learning, the AI tested its predictions of what made a polymer a good contrast agent against a “testing set” of polymers whose effectiveness weren’t labeled, correcting itself when it predicted wrongly and requesting new data. Finally, exploring the chemistry of a representative group of synthesized polymers selected by AI without human supervision, it came up with its own set of rules.
The work was enabled by the robotic system built by the Frank Leibfarth group at UNC Chapel Hill. Leibfarth’s group had built an automated continuous-flow system designed to build polymers that can be used to create plastics, packaging and a number of useful materials. The lab provides the “hands” of the operation. The “brain” was supplied by the Isayev team’s AI. Graduate student Marcus Reis of Leibfarth’s group was the co-first author for the work from UNC.
“Even a small model space leads to intensive computing requirements. Because the calculations are done over a set of classes of machine learning models exhaustively to get maximum information from the data and repeated on each refinement after getting new data, we needed a lot of computing power. [XSEDE] helped us speed up this project,” said Filipp Gusev at CMU.
Through a series of eight refinements, Gusev’s AI was able to narrow a potential 50,000 polymers to a list of only 397 experimentally synthesized. Iterating between the computer and the lab identified the best performing of these candidates. These performed as much as 50% better than current MRI contrast agents.
These winning candidates posed a surprise to the human chemists. Clinical MRI works by detecting changes in a strong magnetic field created by substances in the human body. One family of MRI contrast agents uses the isotope fluorine-19 (19F), which has the ability to interact with dissolved oxygen in body fluids. This interaction can be detected in a strong magnetic field and tells doctors where oxygen is concentrated in living tissues.
Scientists had long thought that more is better in terms of 19F solution concentration – the more 19F atoms that a contrast agent could pack in a smaller space, the better. But 19F also makes the polymer less soluble in water – and if the polymer can’t be dissolved, it can’t be injected.
The leading candidates the AI picked did contain enough 19F to create a strong signal. As they had hoped, the AI had found a “Goldilocks” point of just enough to give a strong signal while still being soluble, a point that humans had not predicted. The result offers hope that AI-guided design can create chemical tools that surpass what human experts can design.
The CMU team reported their results in a paper in the Journal of the American Chemical Society in November, 2021. They would next like to extend the approach toward other types of polymers and organic materials.
O.I. acknowledges support from NSF CHE-1802789 and CHE- 2041108. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number ACI-1548562. Specifically, it used the Bridges-2 system, which is supported by NSF award number ACI-1928147, at the Pittsburgh Supercomputing Center. This research is part of the Frontera computing project at the Texas Advanced Computing Center. Frontera is made possible by the National Science Foundation award OAC-1818253. F.L. acknowledges the UNC Department of Chemistry’s NMR Core Laboratory that provided expertise and instrumentation that enabled this study with support from the National Science Foundation (CHE-1828183 and CHE-0922858).
Source: Ken Chiacchia, Pittsburgh Supercomputing Center




























































