This post was contributed by Pankaj Vats, PhD, and Timothy Harkins, PhD, from NVIDIA Parabricks.
Introduction
Advances in Next Generation Sequencing (NGS) technologies over the last decade have led to many novel discoveries in cancer genomics1,2,3. Whole genome, whole exome and targeted-panel sequencing are more widely used today in clinical research and practice, to diagnose cancers and identify the course of treatment for cancer patients based on tumor mutations. This has led to the formation of consortia dedicated to studying cancer genomics at a large scale3,5. As the volume of data increases, the algorithms and workflows used to analyze the data are proving to be the primary bottleneck. NVIDIA Clara Parabricks addresses this issue by providing an accelerated, scalable, and reproducible software suite optimized for genomic analysis on Graphics Processing Units (GPUs).
In a previous blog post, we discussed the germline variant calling tools available in Clara Parabricks. In this post, we will provide an overview of how to perform somatic variant analysis for cancer workflows using Parabricks from the AWS Marketplace. Somatic variants are genetic alterations which are not inherited but acquired during one’s lifespan, for example those that are present in a tumor. In this post, we will demonstrate how to perform somatic variant calling from matched tumor and normal, as well as tumor-only whole genome and whole exome datasets using an NVIDIA GPU-accelerated Parabricks pipeline and compare the results with baseline CPU-based workflows.
Somatic variant calling pipeline overview
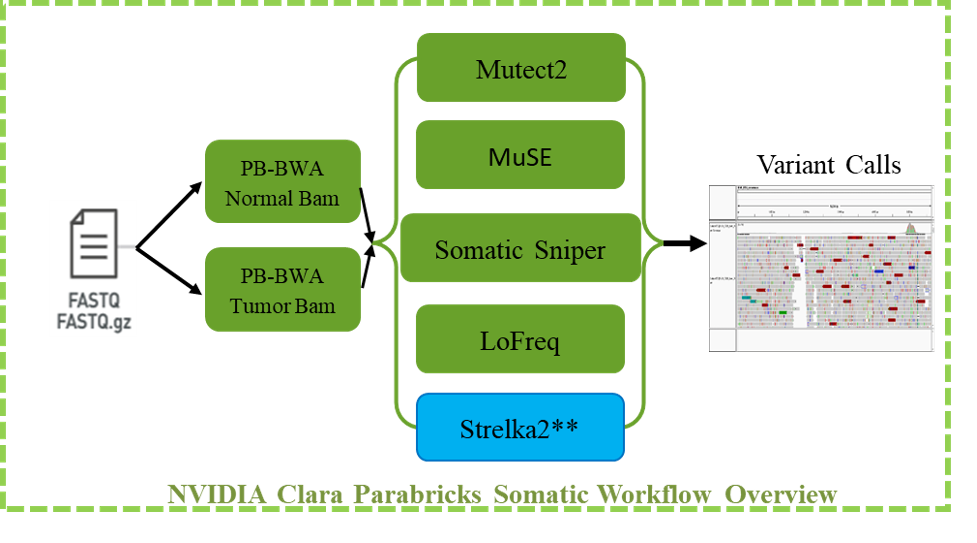
The Parabricks somatic variant workflow starts with tumor and normal bam files as input for five state of art somatic variant calling tools Mutect2, MuSE, Somatic Sniper, LoFreq – which are GPU accelerated in the Clara Parabricks software suite – and Strelka2, which is not. The vcf output of these algorithms can be used to generate consensus vcf or intersection vcf for downstream secondary analysis. Parabricks also provides another important feature where users can provide a set of normal samples to create a panel of normal (PON). Mutect2 can be run in tumor-only mode, with or without a panel of normal (PON).
Dataset used
In this post, we will use the ~50x Tumor/Normal short-read human data from the Somatic Mutation Working Group of the SEQC26-9 consortium. The dataset consists of both whole genome and whole exome HCC1395 (triple negative breast cancer cell line) and HCC1395 BL (matched normal cell line), sequenced on multiple platforms with multiple technical replicates. The WGS data had ~55x mean coverage for tumor and normal, and the WES data had ~100x for tumor and 80x for normal sample. We chose this dataset, because it is well characterized, and the consortium provides a Truth set for the somatic variant calls.
Running NVIDIA Clara Parabricks Somatic Workflow
We detailed how to subscribe and use the NVIDIA Clara Parabricks AMI from the AWS Marketplace in the previous post on benchmarking the germline pipeline. We followed the same prerequisites and initial setup from that post up until “Step 3. Download data” where we shifted to leveraging resources for the somatic pipeline benchmark.
We ran 4 variant callers (Mutect2, Somatic Sniper, Muse and LoFreq) on the SEQC2 dataset. The details of the command line options to download the data and run each of these variant callers can be found at this GitHub repository the Clara Parabricks team have provided for this blog post. You can also find a more detailed overview of somatic variant calling tools available in Clara Parabricks within the NVIDIA documentation.
Results
Runtime comparison with respect to the baseline CPU algorithms versus GPU accelerated NVIDIA Clara Parabricks variant callers.
The NVIDIA Clara Parabricks software can run on different NVIDIA GPUs instances like the Amazon EC2 G4, P3, and P4, but here we will focus on the Parabricks runs on g4dn.12xlarge (4 NVIDIA T4 Tensor Core GPUs) runs in comparison to the baseline callers that were run on an m5.8xlarge (32 vCPU) instance.
Runtime comparison of Parabricks versus the CPU-based baseline callers on seqc2 WGS shows a significant speedup ranging from 4x for LoFreq, 6.5x for MuSE, 9x for Somatic Sniper and 42x acceleration for Mutect2 (tumor/normal mode) relative to the baseline callers detailed runtime for each caller is represented in Figure 2. If users want to run tumor-only somatic variant calling, Parabricks provides the ability to do so with 56x acceleration relative to Mutect2 baseline caller (Figure 2).
Read the full blog to learn more about benchmarking NVIDIA Clara Parabricks Somatic Variant Calling Pipeline on AWS.
Reminder: You can learn a lot from AWS HPC engineers by subscribing to the HPC Tech Short YouTube channel, and following the AWS HPC Blog channel.