Fast sequencing of bacterial and viral strains is critical in identifying public health threats. Data analysis, not sequencing itself, is the challenge with I/O bottlenecks being most problematic. The Public Health England, leveraging DataDirect technology, recently accelerated its sequence process 16x and can now sequence 92 samples in the time previously needed for 12.
PHE was established in 2013 to consolidate specialists from more than 70 organizations into a single public health service. PHE’s MS bioinformatics unit has been involved in the establishment of a Next-Generation Sequencing (NGS) Service that provides the means to sequence the whole genomes of pathogens.
Analysis of pathogen genomes is performed by many groups, but PHE’s Next-Generation Sequencing Processing and Analysis Service will be among the first public health services in the world to offer a routine service for stakeholders to perform whole genome analysis for identification, analysis and characterization of bacteria and viruses, including salmonella, MRSA, HIV and influenza.
To meet stringent demands across the complete research data lifecycle, PHE sought a data management solution that would enable scientists to generate, analyze, archive and share massive amounts of genomics data. In particular, the ability to process multiple whole genome sequencing samples in parallel would enable PHE to meet the service needs of multiple centers, which is necessary to offer a service nationwide; the performance would be especially critical to monitor aggressive pathogens during a major outbreak or emergency response situation.
Working with integrator OCF, PHE selected DDN’s cloud and big data storage platforms, including DDN’s SFA high-performance storage engine in combination with the company’s EXAScaler Lustre File System Appliance and WOS Object Storage platform.
Here’s a diagram showing the storage system and DDN components:
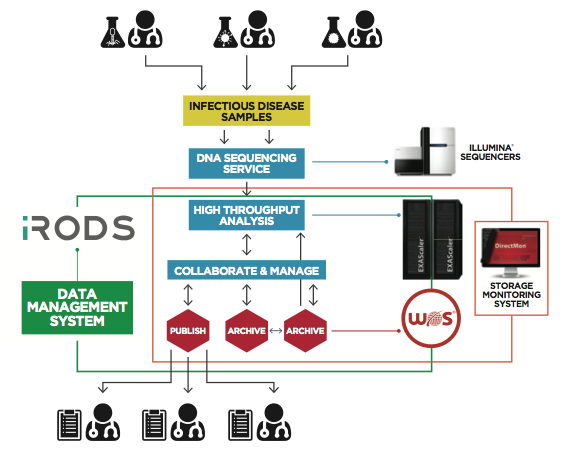 The choice of Lustre as the underlying file system was given careful consideration. PHE was looking for open source software to prevent the technology stack used from being locked into a single vendor, in line with UK government policy. For that reason, Lustre was appealing even though PHE lacked onsite expertise to implement and manage the system. Looking ahead, PHE noted Lustre was being used more by the Top500 supercomputers globally and was felt that this approach would best meet the institution’s current and future needs, despite the learning curve required for deployment and support.
The choice of Lustre as the underlying file system was given careful consideration. PHE was looking for open source software to prevent the technology stack used from being locked into a single vendor, in line with UK government policy. For that reason, Lustre was appealing even though PHE lacked onsite expertise to implement and manage the system. Looking ahead, PHE noted Lustre was being used more by the Top500 supercomputers globally and was felt that this approach would best meet the institution’s current and future needs, despite the learning curve required for deployment and support.
DDN’s ability to integrate with iRODS, which has been gaining traction in life sciences, was also important. It let PHE streamline the data lifecycle for both internal and external scientists who need to ingest, process, analyze, store, share and protect vital genomics research. Key capabilities are:
- With integrated iRODS, researchers now can find data regardless of where it’s physically located by performing an indexed query of user-specified metadata across petabytes of both file and object data.
- The WOS platform will be able to provide an active archive where researchers can access geographically dispersed and replicated file data to make it easy for researchers worldwide to access data long-term for further analysis or publication.
PHE says it is now better positioned to meet the demands of its bioinformatics analysis standards. The team uses BWA, Bowtie and several variant callers for mapping data to reference sequences, variant analysis and mutation detection, which is a key PHE workflow. To perform de novo assembly of new organisms, the team also uses Velvet and Spades. PHE uses Galaxy to provide a web interface for interacting with the HPC cluster and for training purposes. In supporting the scientific lifecycle, PHE also is building a solution that will be able to help people around the world access this data.