In this regular feature, HPCwire highlights newly published research in the high-performance computing community and related domains. From parallel programming to exascale to quantum computing, the details are here.
Modeling urban air pollution using weather, traffic and HPC
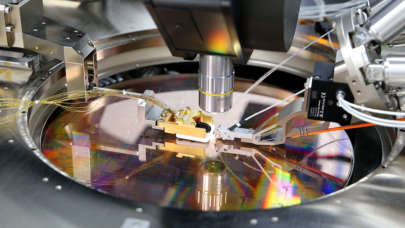
Over three million yearly deaths are at least partially attributable to urban air pollution, of which traffic emissions constitute an enormous portion. These authors, hailing from HLRS and Széchenyi István University in Hungary, conducted a one-year simulation of street-level air pollution, combining a traffic simulation with a computational fluid dynamics simulation and running the coupled model on HPC resources.
Authors: Laszlo Kornyei, Zoltan Horvath, Andreas Ruopp, Akos Kovacs and Bence Liszkai.
Conducting breast histopathology with HPC and deep learning
Breast histopathology (histopathology being the study of tissue diseases) has produced a deluge of data, with these authors from HES-SO Valais-Wallis in Switzerland explaining that “the increasingly intensive collection of [digitized] images of tumor tissue over the last decade made histopathology a demanding application in terms of computational and storage resources.” In their paper, the authors introduce a modular HPC pipeline with three layers for detecting tumor regions in breast lymph node images.
Authors: Mara Graziani, Ivan Eggel, François Deligand, Martin Bobak, Vincent Andrearczyk and Henning Müller.
Performing human-scale brain simulation on a supercomputer
“Unprecedented computational power enables us to build and simulate large-scale neural network models composed of tens of billions of neurons and tens of trillions of synapses,” explain these authors from RIKEN and the University of Electro-Communications in Japan. “Towards this milestone, it is mandatory to introduce high-performance computing technology into neuroscience research.” In their article, they review simulation studies specific to the cerebellum, presenting results of a recent simulation of a human-scale cerebellar network model composed of 86 billion neurons that was run on the (now retired) supercomputer K.
Authors: Tadashi Yamazaki, Jun Igarashi and Hiroshi Yamaura.
 Reviewing current and future converted cloud-HPC workflows at LLNL
Reviewing current and future converted cloud-HPC workflows at LLNL
For these researchers from Lawrence Livermore National Laboratory (LLNL), scientific workflows “require the integration of cloud technologies with traditional HPC to make discoveries.” In their paper, they present trends in these converged workflows and remaining gaps at LLNL, describing successful workflow patterns and highlighting the techniques they are applying to address ongoing challenges at the lab.
Authors: Daniel J. Milroy, Stephen Herbein and Dong H. Ahn.
Annotating HPC metadata in the age of dark data
“Dark data” – data acquired, but unused – is looming over fields with massive data collection apparatuses, worsened and enabled by missing metadata that could make the data functional for an organization. In this paper, Björn Schembera of HLRS presents ExtractIng, a “generic automated metadata extraction toolkit” aimed at automatically assigning scattered metadata to simulation files in a uniform manner and ameliorating dark data in HPC.
Author: Björn Schembera.
 Fostering remote visualization for two HPC sites
Fostering remote visualization for two HPC sites
“Remote visualization is of crucial importance to access infrastructure, data and computational resources and, to avoid data movement from where data is produced and to where data will be analyzed,” say these authors, who hail from the Industrial University of Santander and Oak Ridge National Laboratory. In the paper, they present two approaches deployed by two HPC centers: the SC3 center in Colombia and the Oak Ridge Leadership Computing Facility. They summarize experiences, technologies, use cases and challenges among the two cases.
Authors: César A. Bernal, Carlos J. Barrios and Benjamín Hernández.
Running predictive analytics on genomic data with HPC
Next-generation sequencing (NGS) technologies, write these authors from Canada and Nigeria, “have led to tremendous reduction in sequencing time and given rise to the production and collection of high volumes of genomic datasets.” Predicting protein-coding genes – a valuable function for protein synthesis and other tasks – is computationally intensive. The authors use their paper to explore predictive analytics for genomic data, presenting a “scalable naïve Bayes-based algorithm that is deployed over a cluster of Apache Spark framework for efficient prediction of genes in the genome of eukaryotic organisms.”
Authors: Carson K. Leung, Oluwafemi A. Sarumi and Christine Y. Zhang.
Do you know about research that should be included in next month’s list? If so, send us an email at [email protected]. We look forward to hearing from you.