SCIENCE & ENGINEERING NEWS
Ithaca, N.Y. — Gail Robinson reports that Cornell University researchers hope to develop a much faster way to determine the shape of protein molecules by analyzing X-ray data on a computer.
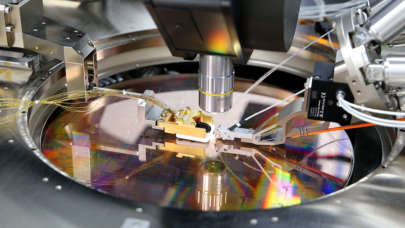
When a beam of X-rays is fired through a crystallized protein sample, the beam is scattered into a pattern that depends on the arrangement of atoms in the crystal. By decoding that pattern, experts can find the arrangement of the atoms and the shape of the protein molecule. But the decoding process used to find the structure of a large protein can take weeks or even months.
Cornell researcher Veit Elser has received a three-year National Science Foundation grant of $234,320 to develop new computer algorithms for X-ray crystallography. The grant comes from the agency’s Information Technology Research initiative.
Crystallographers measure the amplitude of the X-rays emerging from a sample by scanning across a plane on the output side to deduce the shape of the object under study. But amplitude is only half the information contained in a stream of X-rays. Just as important is the phase of the photons in the beam.
So far, X-ray crystallographers don’t have good ways to measure phase. The new approach simply reanalyzes the data using a mathematical trick. By a quirk of the math, out of the enormous number of phase patterns possible, there’s only one that will combine correctly with one particular pattern of amplitudes. But hitting on that one pattern is the problem. An exhaustive computation of all possibilities would take the most advanced supercomputer a time equal to the age of the universe to try out every possibility.
The technique uses a simple test to see if the computer is converging on a solution. The hypothetical structure that a given combination of phases and amplitudes predicts must have the same total charge as the number of electrons in the real protein. If the hypothetical charge doesn’t add up, the algorithm immediately abandons the calculation and searches for a new combination. The hope is that the new algorithm will be able to reconstruct the protein molecule configuration from current X-ray diffraction experiments in a matter of minutes.
============================================================