Leading supercomputer vendor Cray and datacenter/cloud provider the Markley Group today announced plans to jointly deliver supercomputing as a service. The initial offering provides access to Cray’s Urika GX platform, housed in Markley’s massive Boston datacenter, and focused on the many biotechs in the region. The partners say the service is unique and they plan to address other verticals with a range of Cray products over time.
“We want to take a targeted approach and are going to be really thoughtful about what vertical is next or what type of infrastructure best solves the use case represented by that vertical and where the need or demand is,” said Fred Kohout, senior vice president of products and chief marketing officer, Cray. “We want to be customer-led here.” Certainly the Boston-Cambridge area is a mecca for large and small life sciences organizations in both industry and academia.

Supercomputing as a service, say Cray and Markley, will make supercomputing available to many users who are unable to afford or support such resources themselves or who only need those resources sporadically. They also argue supercomputing produces a significant performance advantage over traditional HPC clusters, in this case on the order 5X for the genomics workloads evaluated so far.
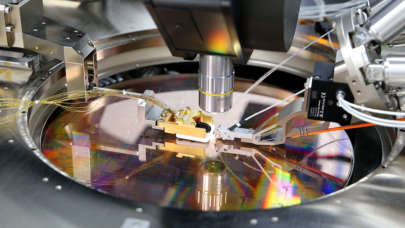
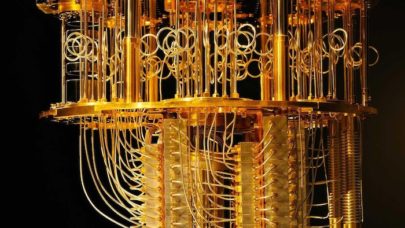
“This is supercomputing. It sounds like a marketing term but it’s really different. We are not talking about 1000 Dell blades all in the same datacenter. We are talking about the Cray Aries interconnect and optimizations such Cray Graph Engine (CGE) that are qualitatively different than just having lots of CPUs close to each other,” said Patrick Gilmore, chief technology officer, Markley.
No doubt there will be kinks to iron out, but supercomputing as a service is an interesting paradigm shift and potential market expander for Cray. The partners declined to say much about pricing, other than they were looking at prices for HPC-in-the-cloud resources and that there would be a premium over those; how much wasn’t revealed. It will be more than 10 percent higher but it won’t “be a discouraging” premium say the partners.
Cray bills the Urika GX as the first agile analytics platform that fuses supercomputing abilities with open enterprise standards. Its Cray Graph Engine provides optimized pattern-matching and is tuned to leverage the scalable parallelization and performance of the Urika-GX. These strengths are particularly valuable for many bioinformatics tasks.
“Research and development, particularly within life sciences, biotech and pharmaceutical companies, is increasingly data driven. Advances in genome sequencing technology mean that the sheer volume of data and analysis continues to strain legacy infrastructures,” said Chris Dwan, who led research computing at both the Broad Institute and the New York Genome Center. “The shortest path to breakthroughs in medicine is to put the very best technologies in the hands of the researchers, on their own schedule. Combining the strengths of Cray and Markley into supercomputing as a service does exactly that.”
As explained by Jeff Flanagan, executive vice president, Markley Group, “the service will not be offered as a ‘partition service’ but as a reservation service. Companies and institutions will have the opportunity to reserve time on the various Cray machines and we are starting with the Urika GX.” One attractive aspect of starting with the Urika GX is that it looks a like a lot like standard Linux box and has a fair amount of pre-installed software (Hadoop is one example), according to Ted Slater, global head of healthcare and life sciences at Cray.
Markley and Cray have tried to remove much of the heavy lifting required for running finicky supercomputers; still, using the service isn’t trivial. As part of the pre-staging, users need to move their data into the Markley datacenter and also make sure their software will actually run when the user’s schedule time on the Urika GX occurs. This can take some time (e.g. a month). That said Markley’s datacenter has a variety of high performance resources (InfiniBand/100Gig Ethernet, petabytes of object storage, fast SSDs, etc.).
Uploading data shouldn’t be problem for most clients says Gilmore, “If you are in New England the Markley datacenter is pretty much the center of the Internet – something like 70 percent of the internet fiber goes through that building so there are lots of ways to get into the building. We’ll get you a connection either a VPN or a direct fiber. Most of the genomics customers we’re talking to are already customers with colocation [here], so there is probably direct fiber from their offices, their sequencers, to the building.”
Data can live on a storage array users already have in the colocation facility or be placed on a Markley array. Cray and Markley have also set up a virtualized version of the Urika GX to serve as a test platform for scripts, and “to make sure they don’t waste time on a very, very expensive, very fast supercomputer, when the reservation comes up.” Markley will then preload the data or make the connection to their array depending upon their preference.
“We are actually going to put the virtual machine behind a load balancer so that when you log onto it and test it, when it’s your turn to come up, you’ll just shut down the virtual machine. We will migrate the virtual machine onto the Cray for you. We’ll reconfigure the load balancing so you don’t actually have to do anything different. It works just like it did when on the virtual machine. We’ll also be transferring all the data for you,” said Gilmore. “So there is a little bit of preorganization to do, but the idea is to make this is as easy and seamless to do as possible. Users log on again, make sure the program works properly, then Monday morning (at the prearranged reserved time) they can turn on the jobs and away they go.”
Cray and Markley say little about the actual projects they did the beta testing on; however on the Cray web site there is brief account of genomics project conducted by the Broad Institute using the Urika GX. It no doubt has lessons. Hail, an open source scalable framework for exploring and analyzing genetic data at massive scale, was used in the Broad project.
Hail is built on top of Apache Spark, and can analyze terabyte-scale genetic data. “Still under active development, Hail is used in medical and population genomics at the Broad for a variety of diseases. It also serves as the core analysis platform for the Genome Aggregation Database (gnomAD) – the largest publicly available collection of human DNA sequencing data, and a critical resource for the interpretation of disease-causing genetic changes,” according to the Cray document.
Hail is also the tool that’s pre-installed on the Cray-Markley Urika GX offering although Slate says users can choose to port the tool of choice, such as GATK (also a Broad project). Slater said Cray has also been working with a genome assembler on its XC platform and is working to port it to GX for evaluation.